Bishal G. Tamang, Ph.D.
Molecular Crop Physiologist
Ainsworth Lab ·
Institute for Genomic Biology
University of Illinois Urbana-Champaign

About
I am a Research Scientist in the Ainsworth Lab at the University of Illinois, working within the RIPE project at the Institute for Genomic Biology. My research program integrates genetics, remote sensing, and computational biology to improve soybean canopy architecture for greater photosynthetic efficiency and yield.
I developed over 200 backcross lines and CRISPR-edited soybean targeting the narrow leaf (ln) locus to optimize leaf area index. I combine UAV and satellite imagery with machine learning to predict plant traits at scale, and use these predicted phenotypes for QTL mapping and phenomic selection. My earlier work characterized transcriptomic responses to flooding and drought stress in soybean, and transpiration dynamics in cereal crops.
Research Areas
- Soybean canopy architecture & leaf shape genetics
- 3D canopy modeling & radiation-use efficiency
- UAV and satellite remote sensing for trait prediction
- Machine learning for genomic-assisted breeding
- QTL mapping with predicted phenotypes
- Transcriptomics & gene function (GmJAG1/ln)
- Plant stress physiology (flooding, drought, transpiration)
Research Focus
Understanding and improving crop productivity through physiological and molecular approaches.
Canopy Architecture & the ln Locus
Developed 200+ BC3 near-isogenic soybean lines and CRISPR-edited GmJAG1 knockouts to dissect how narrow leaf shape reduces leaf area index and improves light distribution through the canopy. Two U.S. provisional patents filed.
Read in Plant Physiology →Remote Sensing & Machine Learning
Using multispectral UAV and satellite (Sentinel-2, Planet) imagery with XGBoost, random forest, and SVM models to predict peak LAI and yield across multi-environment soybean trials with hundreds of genotypes.
Manuscript in preparationQTL Mapping with Predicted Traits
Applying mixed-model GWAS to ML-predicted BLUPs from remote sensing data to identify genetic loci for canopy traits. Identified 70 significant SNPs (29 independent signals after LD pruning) replicated across field locations.
Manuscript in preparationPhenomic Selection
Integrating spectral reflectance data into genomic prediction frameworks. Satellite-derived phenomic selection achieves prediction accuracies comparable to UAV platforms, enabling scalable breeding applications.
Manuscript in preparationTranscriptomics & Gene Function
RNA-seq and single-cell transcriptomic analysis of GmJAG1, the narrow leaf gene. JAG1 acts as a transcriptional repressor of cell cycle regulators including D-type cyclins and histone deacetylases in leaf primordia.
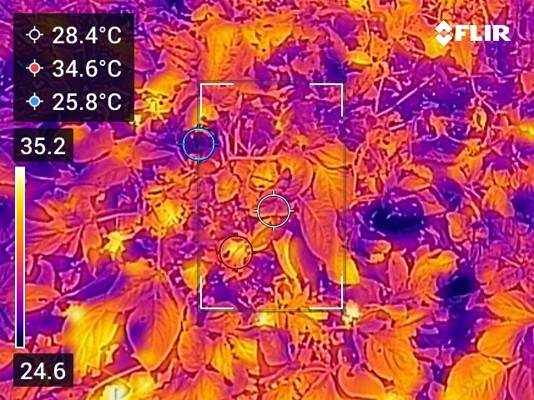
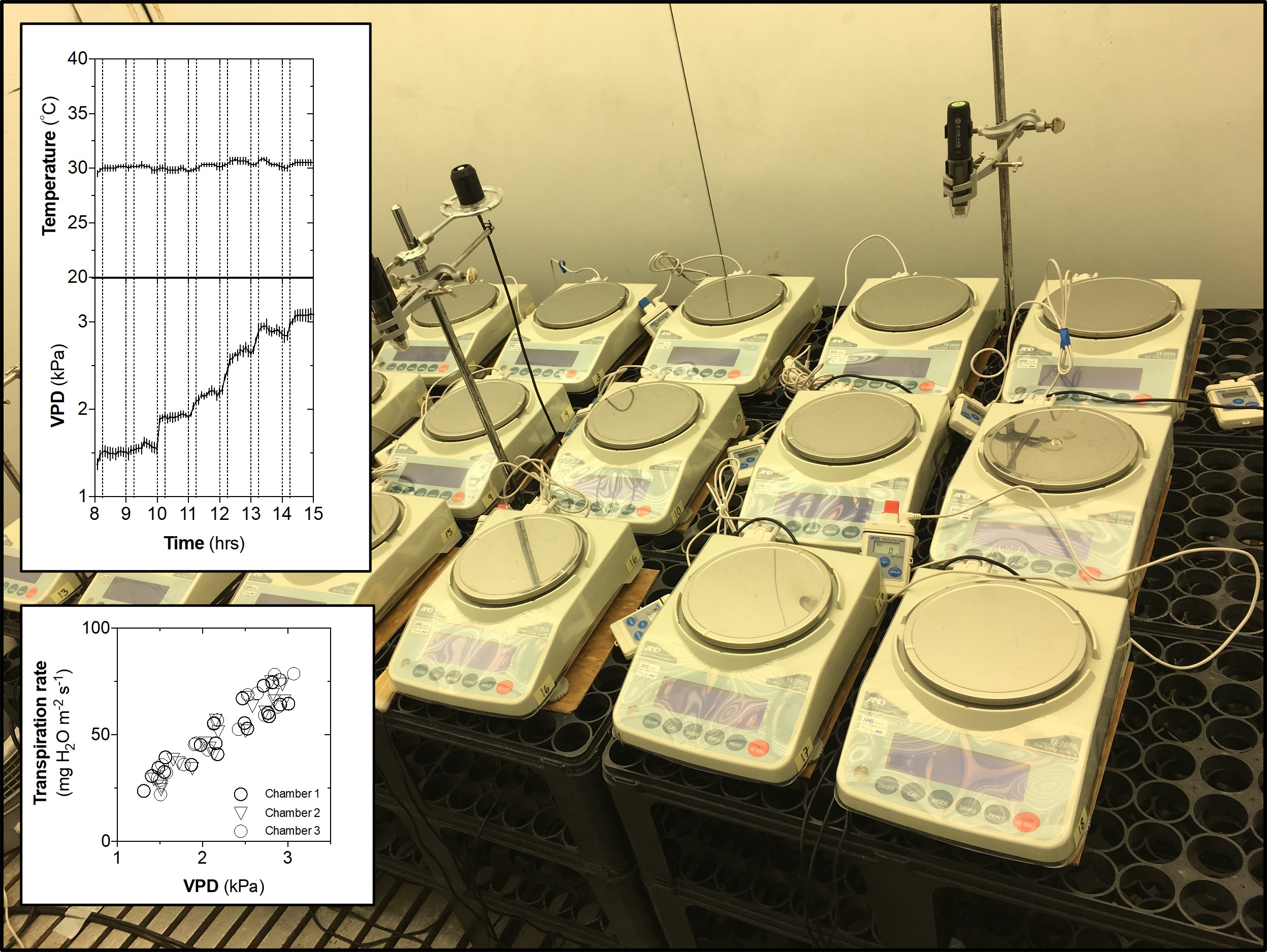
Read preprint on bioRxiv →Stress Physiology & Transpiration
Characterized overlapping and distinct transcriptomic and hormonal responses to flooding and drought in soybean. Earlier work on nocturnal transpiration dynamics and circadian control of water use in cereal crops.
Read in The Plant Journal →3D Canopy Modeling
3D ray-tracing of GmJAG1 near-isogenic canopies shows that narrow-leaflet plants achieve ~16% higher canopy CO2 assimilation than broad-leaf siblings despite ~28% lower leaf area, by distributing absorbed light across more individual leaflets and shifting more leaves into the high-quantum-yield region of the photosynthesis–PAR curve.
Manuscript in preparationSame leaves, different shape, more carbon
In soybean GmJAG1 near-isogenic lines, narrow-leaflet plants have ~28% less total leaf area than their broad-leaf sister lines — yet our 3D canopy simulations show ~16% higher canopy CO2 assimilation in the narrow line at peak season.
The puzzle: both canopies absorb essentially the same fraction (~92%) of incoming PAR. The advantage doesn't come from intercepting more light.
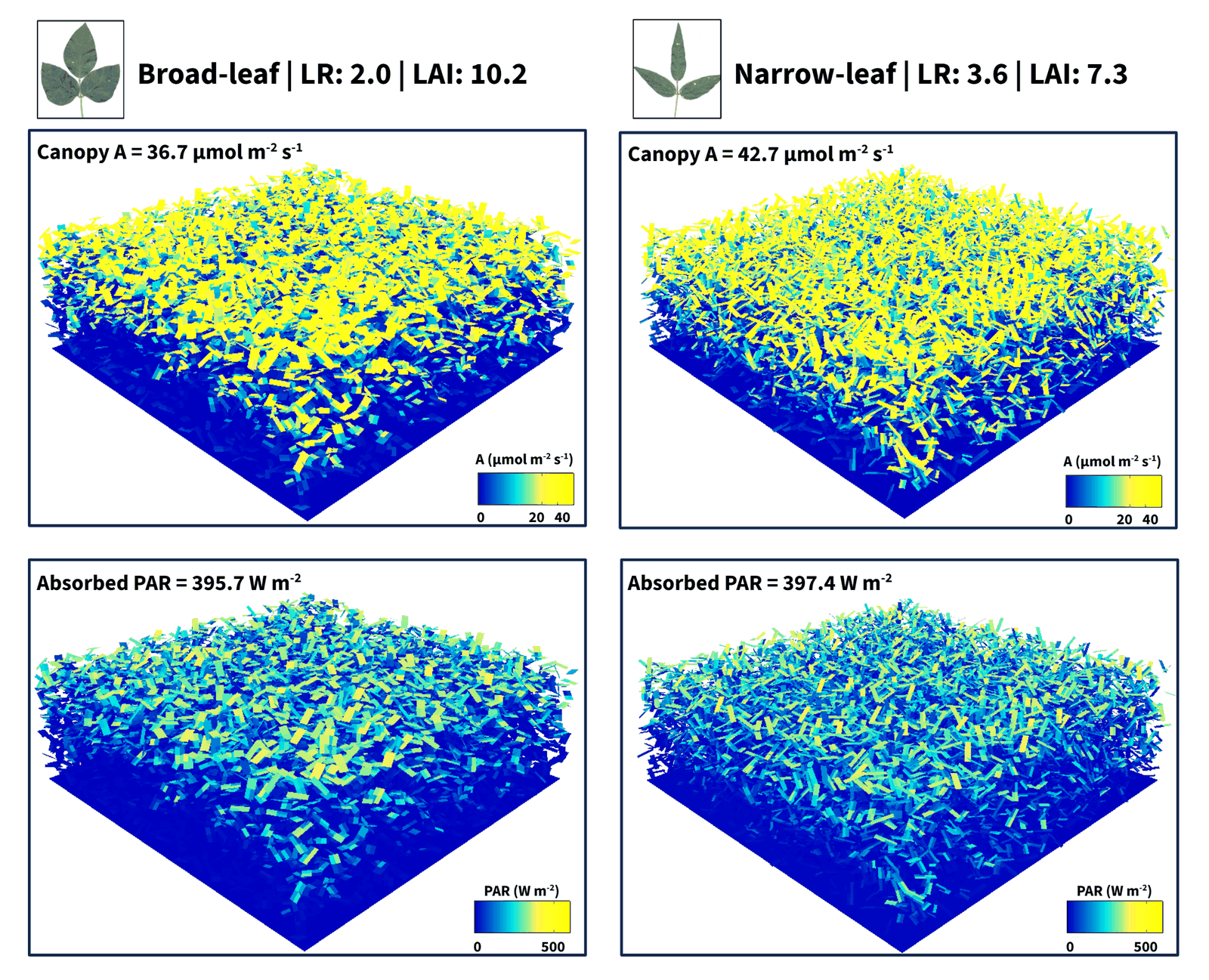
It comes from how that light is distributed across leaflets. Narrow leaflets at matched count push more individual leaves into the high-quantum-yield region of the photosynthesis–PAR curve, with fewer top leaves saturated and fewer bottom leaves in deep shade.
Net result: better per-photon carbon conversion at the canopy scale — the conversion-efficiency leg of Monteith's productivity framework.
The implication for breeding: leaf architecture, not just leaf area, drives canopy radiation-use efficiency. Two NILs with the same leaflet count and the same plant density can have meaningfully different canopy carbon flux based on a single morphological gene.
Scientific Contributions
For a complete list, visit my Google Scholar profile.
Breeding soybean for reduced leaf area index
Inventors: Tamang BG, BW Diers, EA Ainsworth
Application No.: U.S. Provisional 63/781,061
Filed: March 31, 2025
Soybean with stacked genes to improve photosynthesis
Inventors: Tamang BG, BW Diers, EA Ainsworth
Application No.: U.S. Provisional 63/781,059
Filed: March 31, 2025
-
Registration of four soybean lines in a common genetic background with contrasting leaf shapeJournal of Plant Registrations, 2025
-
Bigger is not always better: optimizing leaf area index with narrow leaf shape in soybeanPlant Physiology, 2025
-
Association between xylem vasculature size and freezing survival in winter barleyJournal of Agronomy and Crop Science, 2021
-
Overlapping and stress-specific transcriptomic and hormonal responses to flooding and drought in soybeanThe Plant Journal, 2021
-
Sheathing the blade: Significant contribution of sheaths to daytime and nighttime gas exchange in a grass cropPlant, Cell and Environment, 2020
-
Variability in temperature-independent transpiration responses to evaporative demand correlate with nighttime water usePlanta, 2019
-
Nightly business: Links between daytime canopy conductance, nocturnal transpiration and its circadian controlEnvironmental and Experimental Botany, 2018
-
Plant Adaptation to Multiple Stresses during Submergence and Following DesubmergenceInternational Journal of Molecular Sciences, 2015
-
Physiological and transcriptomic characterization of submergence and reoxygenation responses in soybean seedlingsPlant, Cell and Environment, 2014
News & Highlights
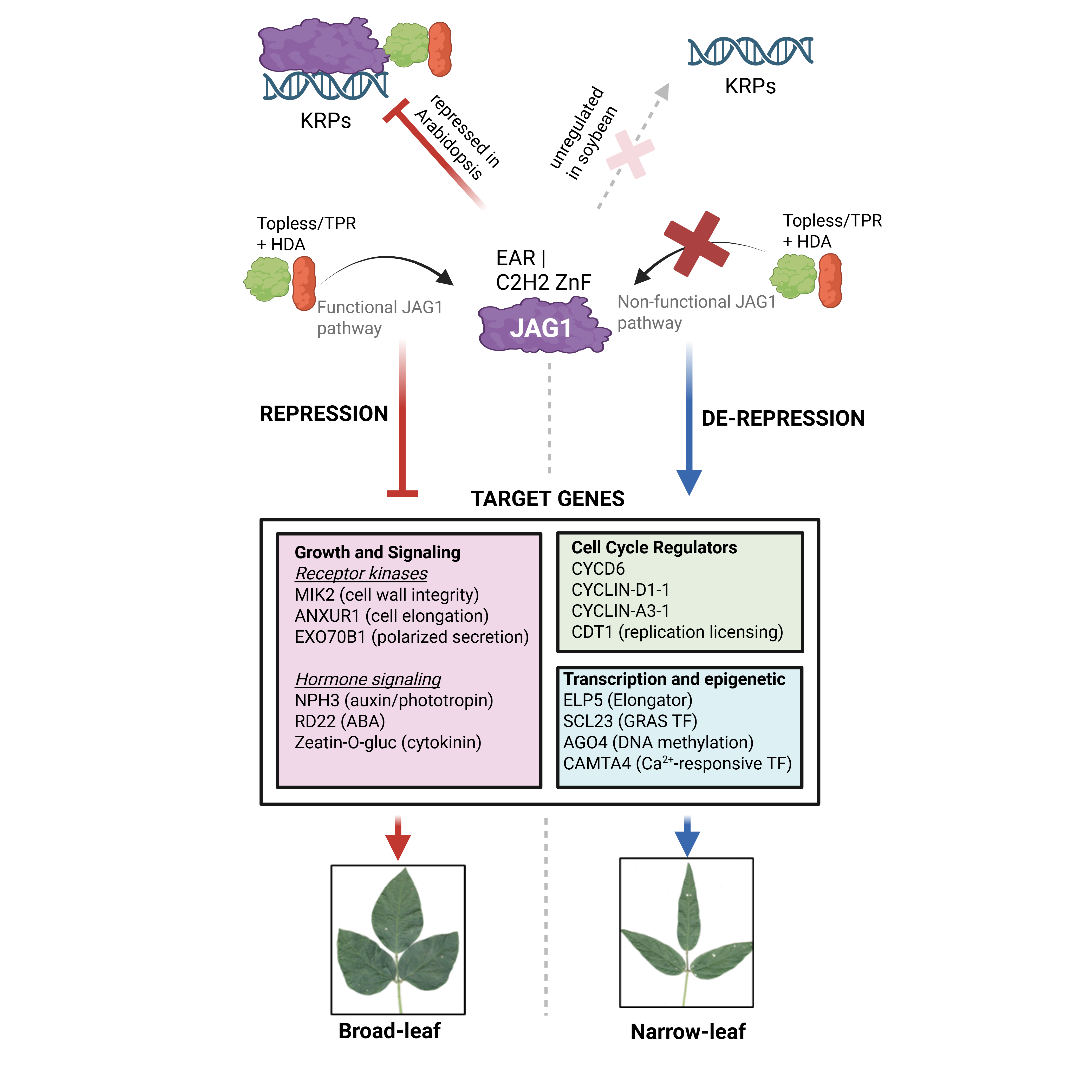
New Preprint on bioRxiv
Our preprint on the genome-wide transcriptional targets of GmJAG1, the gene behind the narrow leaflet trait in soybean, is now available on bioRxiv.
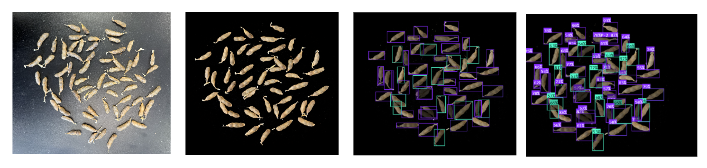
Deep Learning Pipeline for Soybean Pod Phenotyping
Building a deep learning pipeline to segment and count soybean pods from scanned images, enabling high-throughput phenotyping of yield components for the NSPP population.

New Publication in Journal of Plant Registrations
Our paper on the registration of four soybean germplasms with contrasting leaf shapes in a common genetic background is now published.

New Publication in Plant Physiology
Our paper "Bigger is not always better: optimizing leaf area index with narrow leaf shape in soybean" is now published in Plant Physiology!

FAO Global Agrifood Biotechnologies Conference
Presented our research on soybean canopy architecture optimization at the prestigious FAO conference in Rome, Italy.

Speaker at AGBT Agricultural Meeting
Speaker at the Advances in Genome Biology and Technology (AGBT) Agricultural Meeting in Orlando, FL.

New Publication in Functional Plant Biology
Our paper on high-throughput phenotyping of soybean transpiration response curves is now published.
RIPE Project Ambassador
Honored to be selected as a RIPE project ambassador for 2025-2026, promoting our mission to improve global food security.

Soybean Breeders Workshop Presentation
Presented findings on CRISPR-edited soybean lines with altered canopy architecture at the annual workshop in St. Louis.
Research in Action
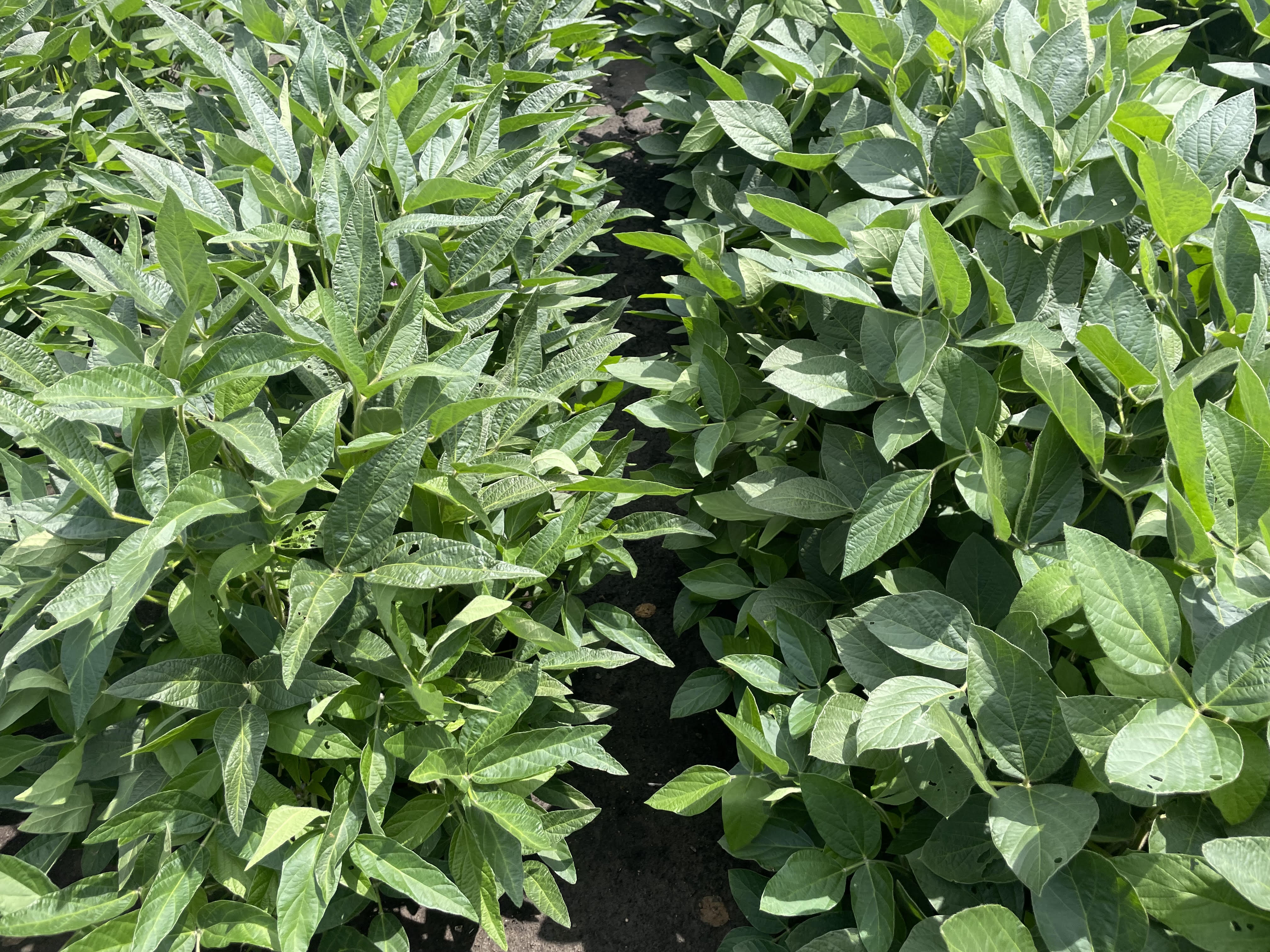
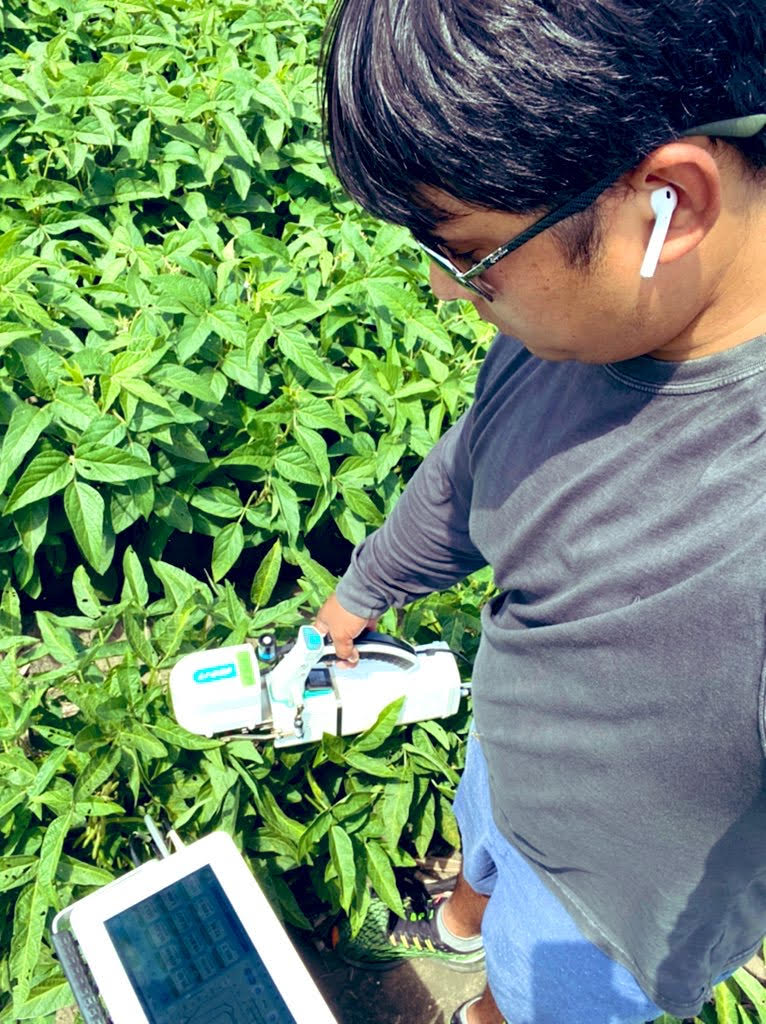
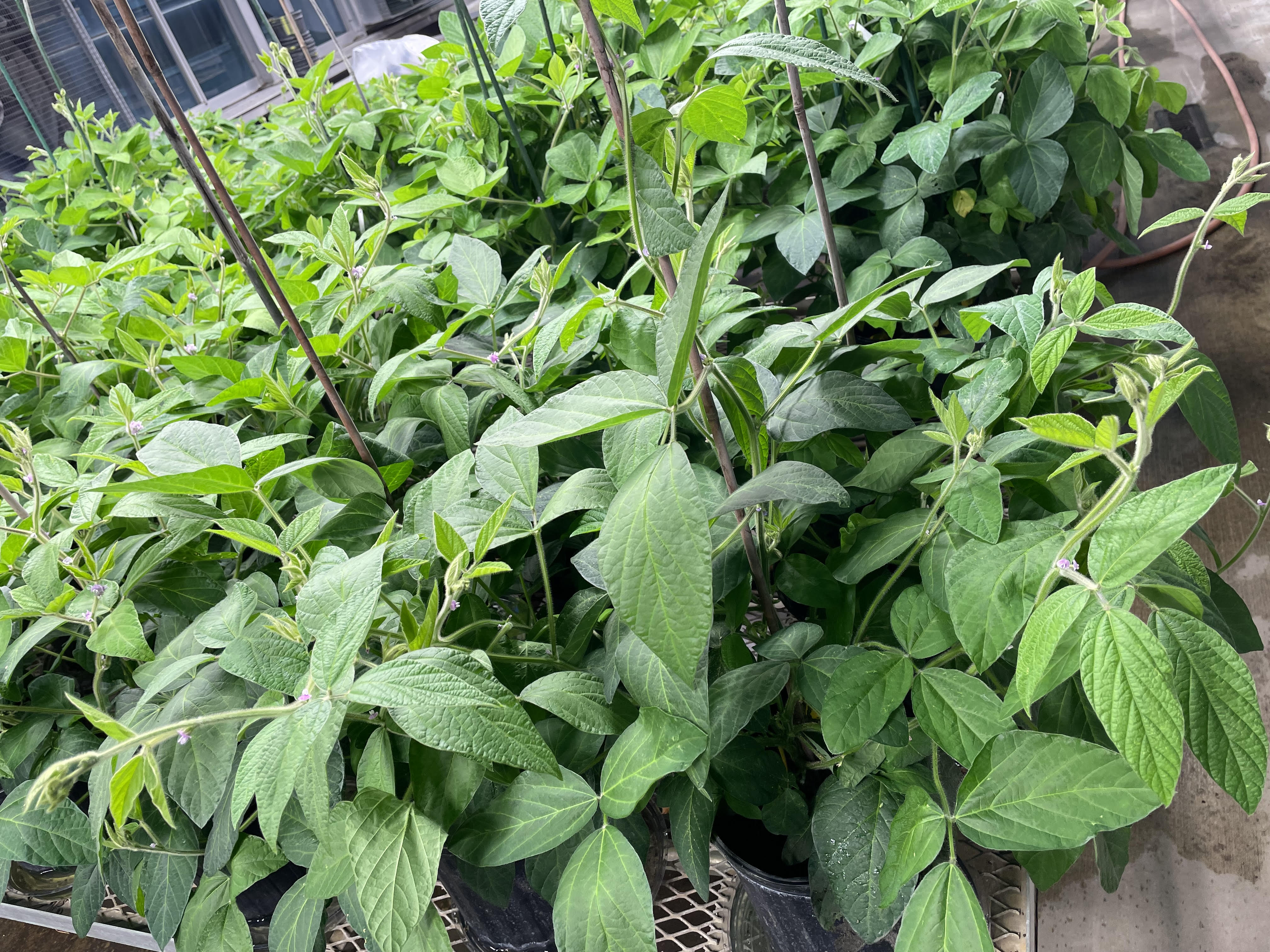
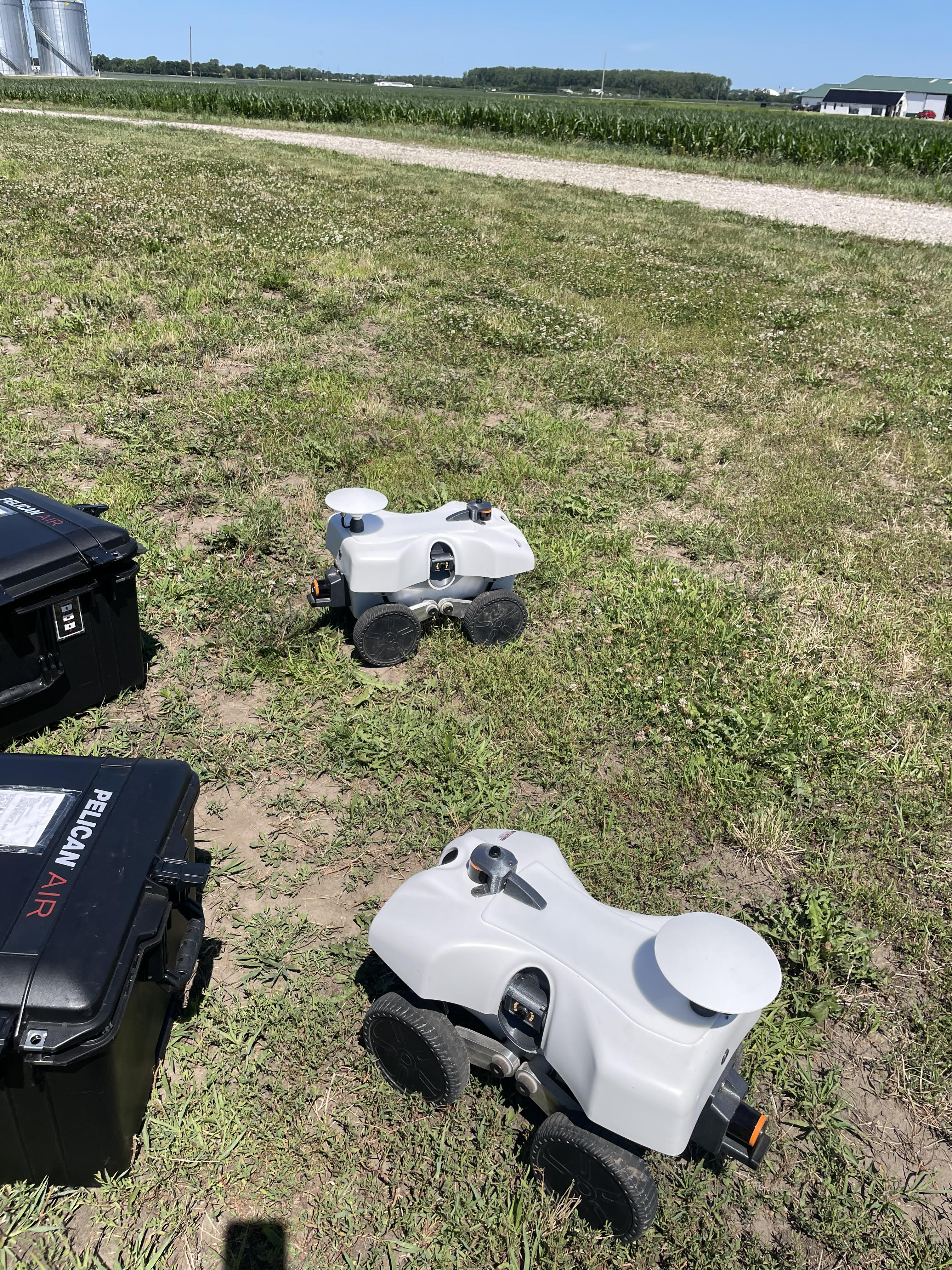
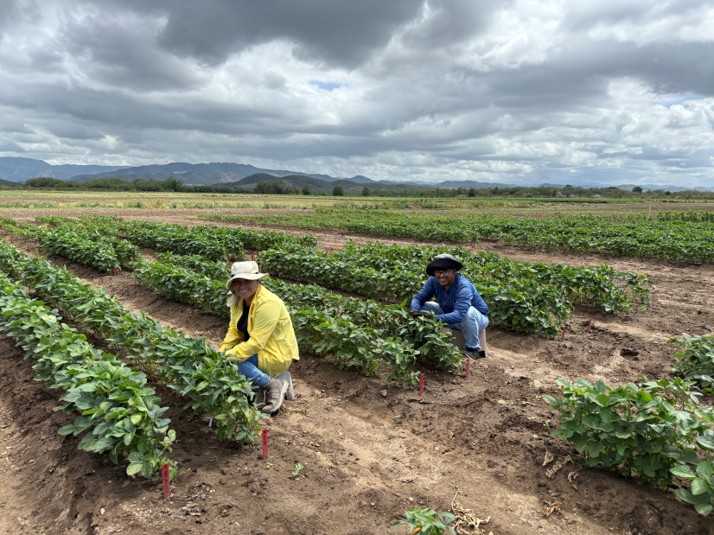
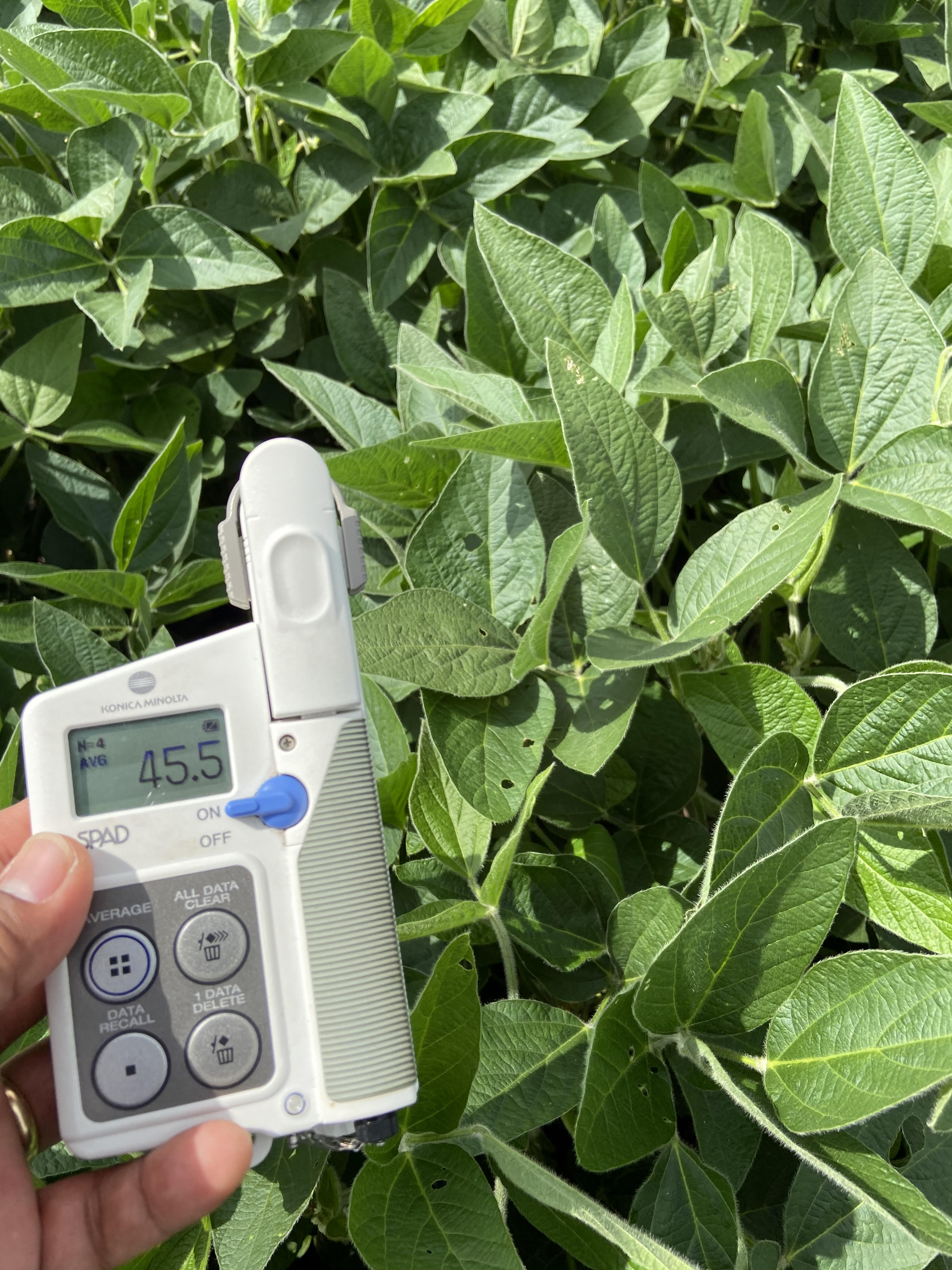
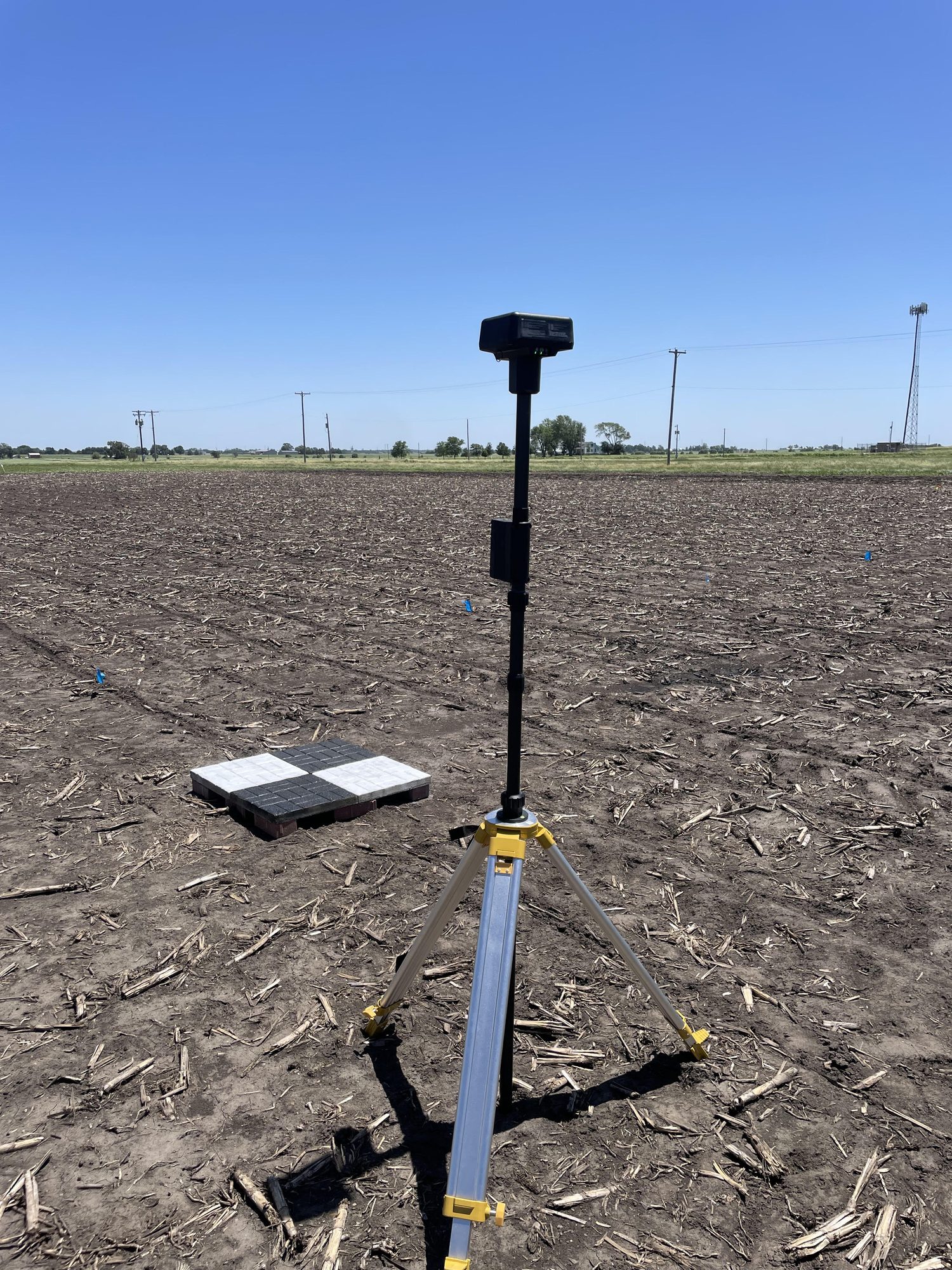
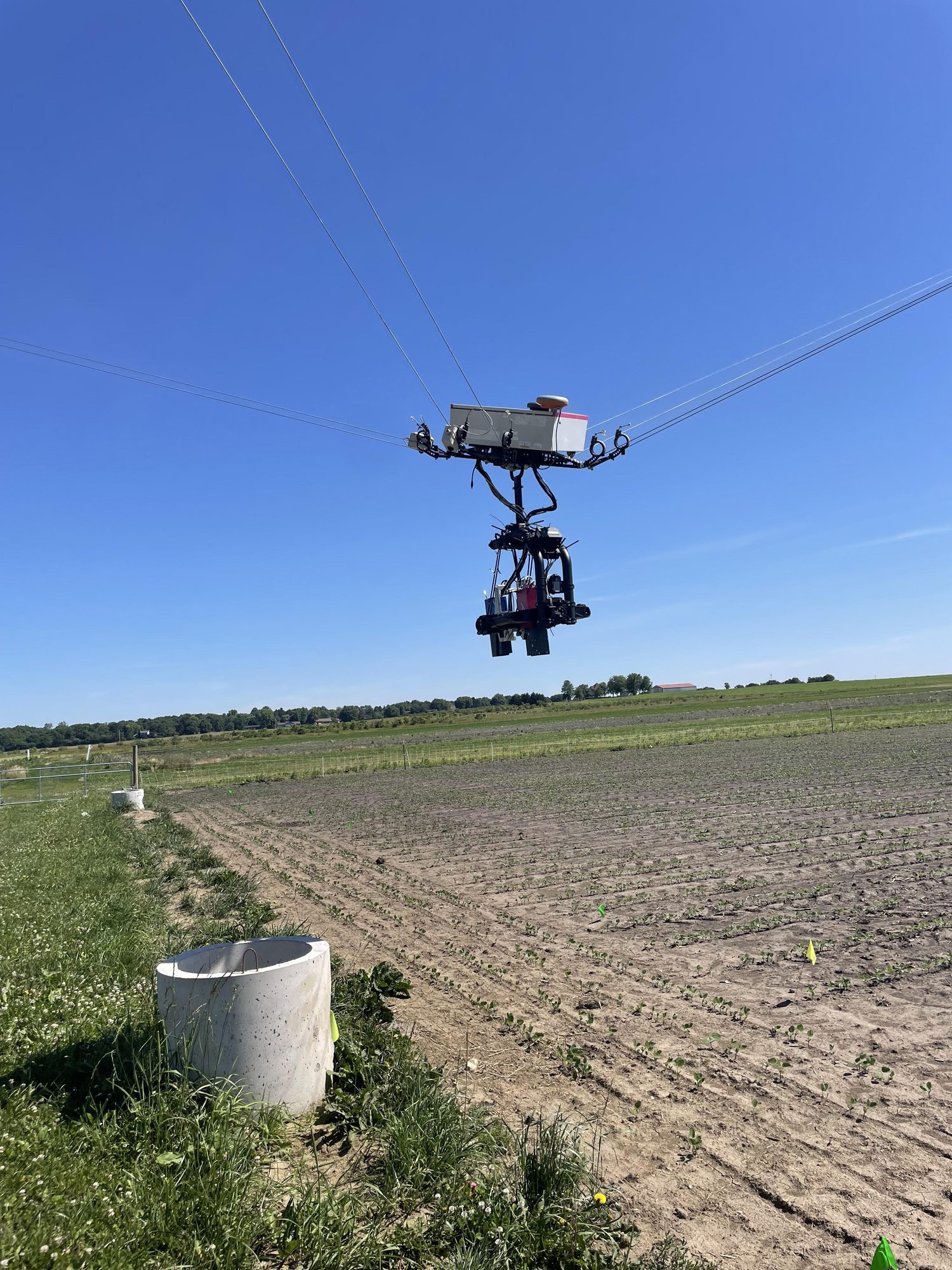
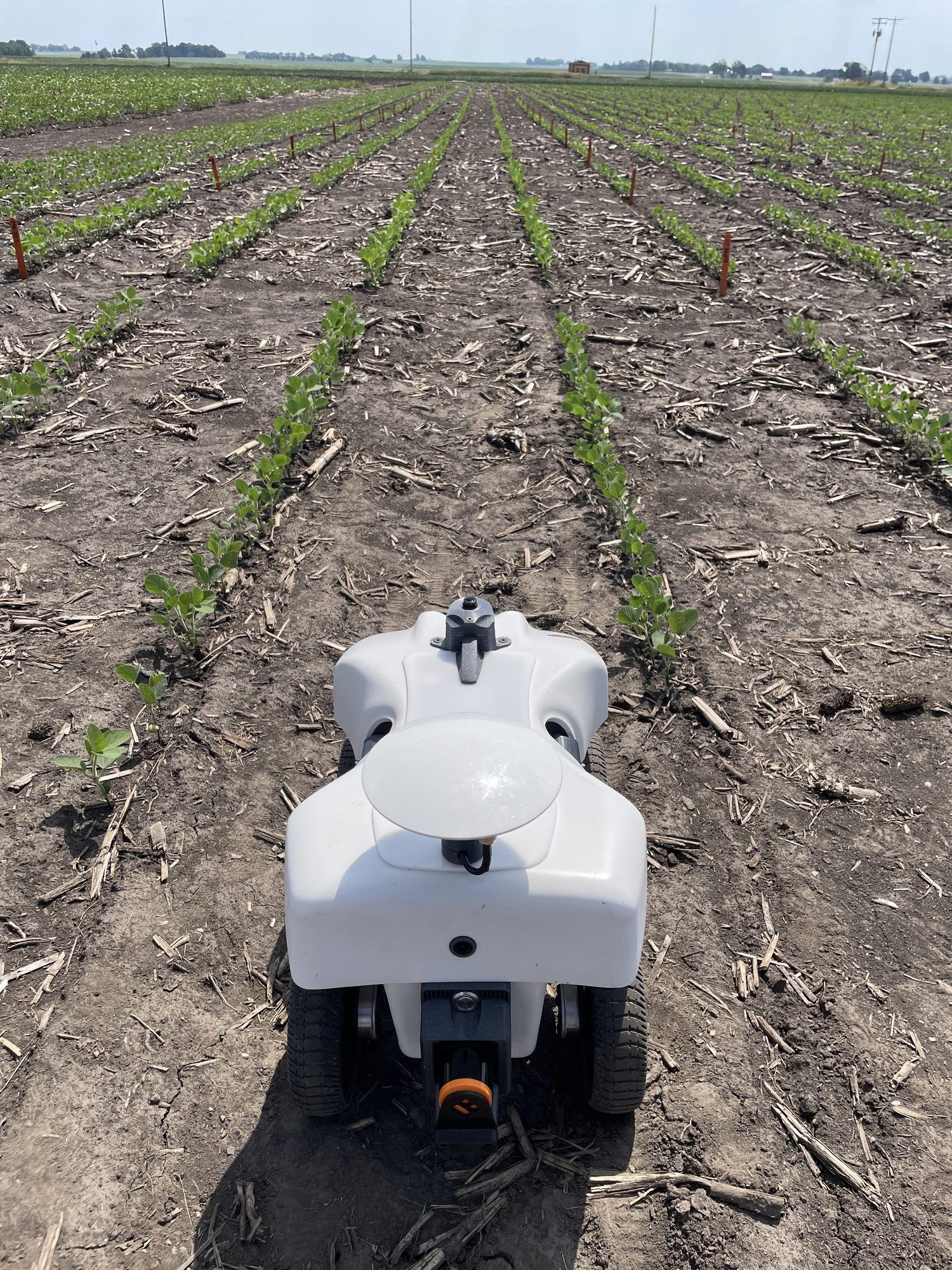
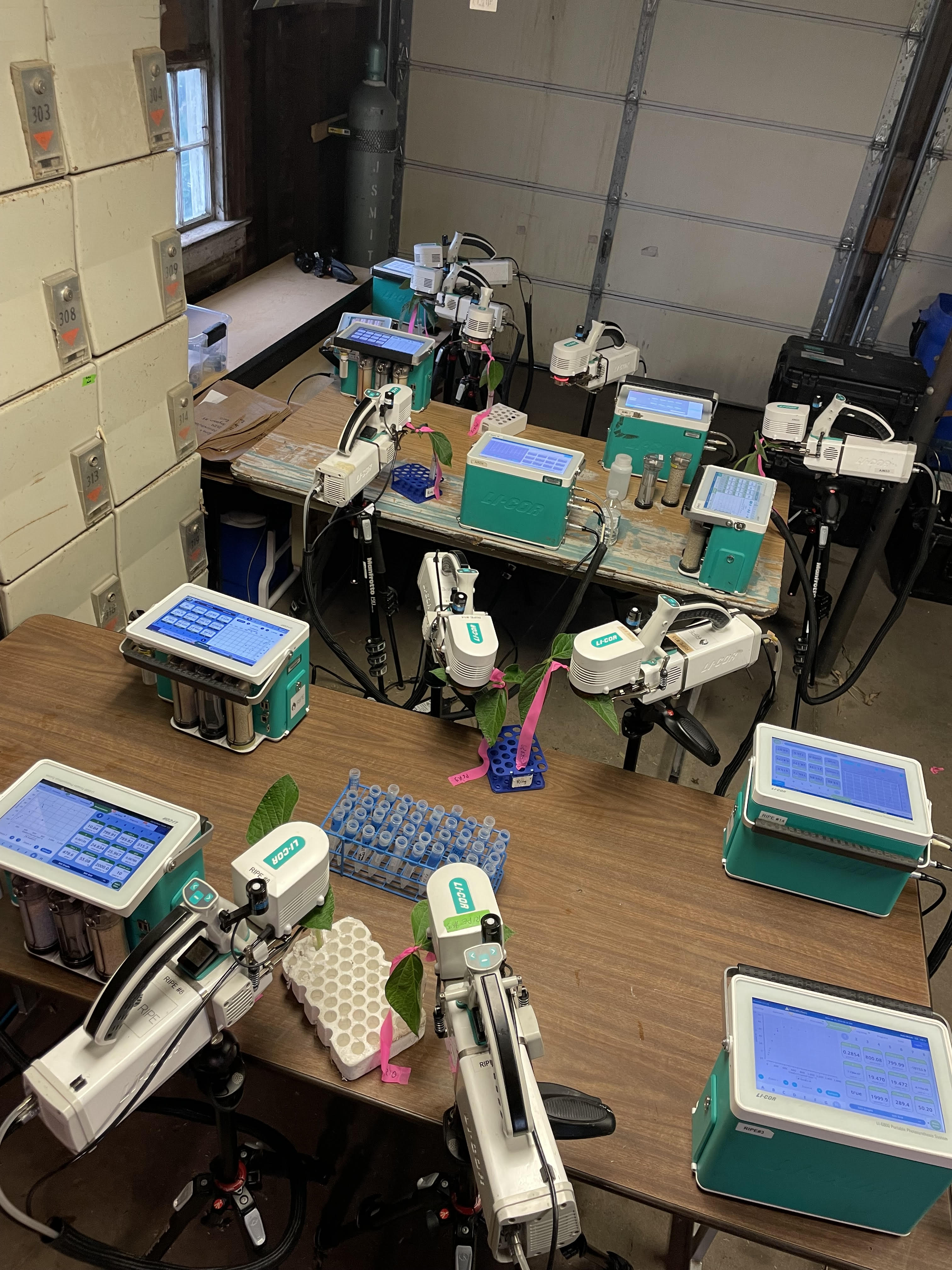
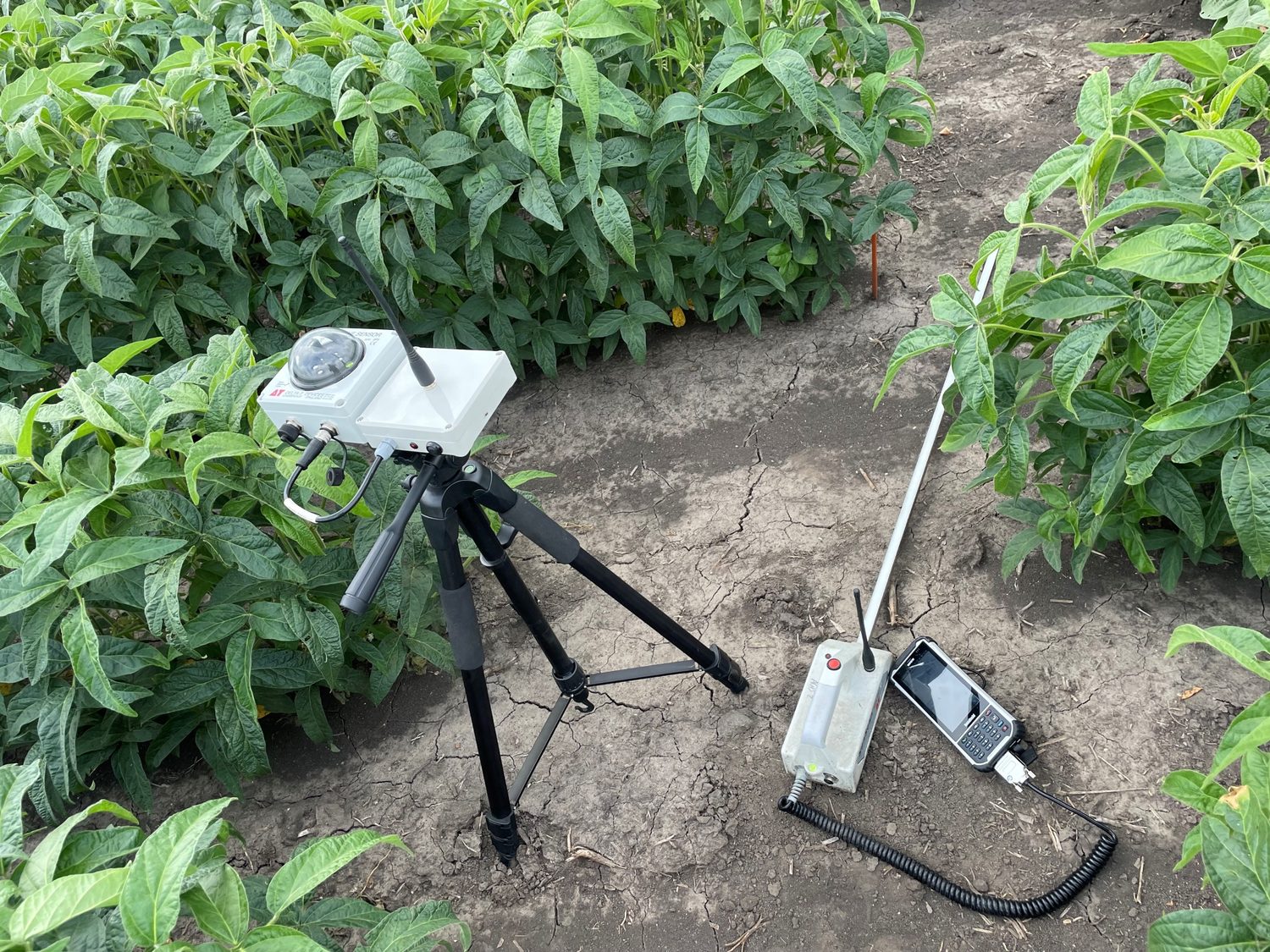
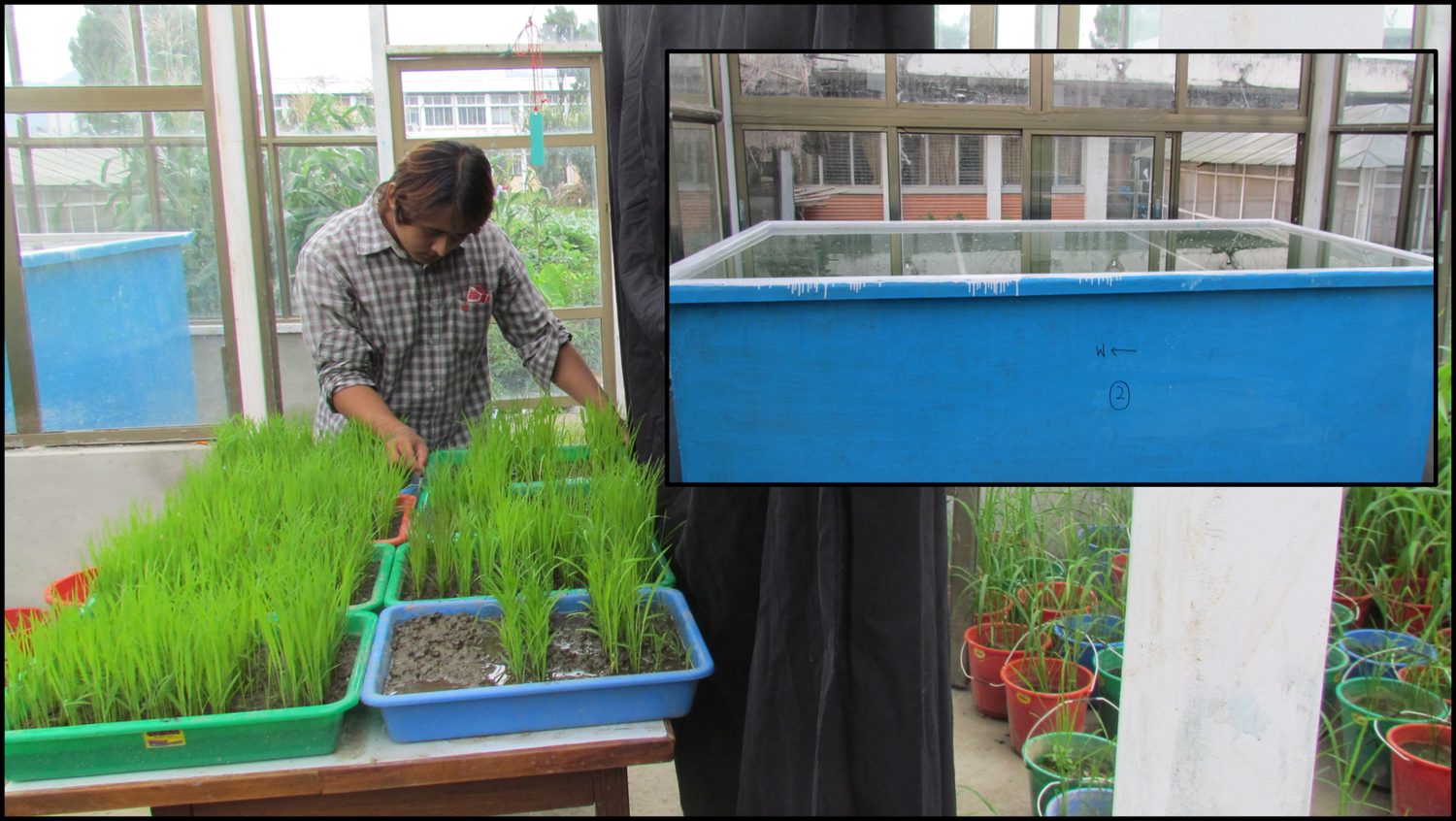
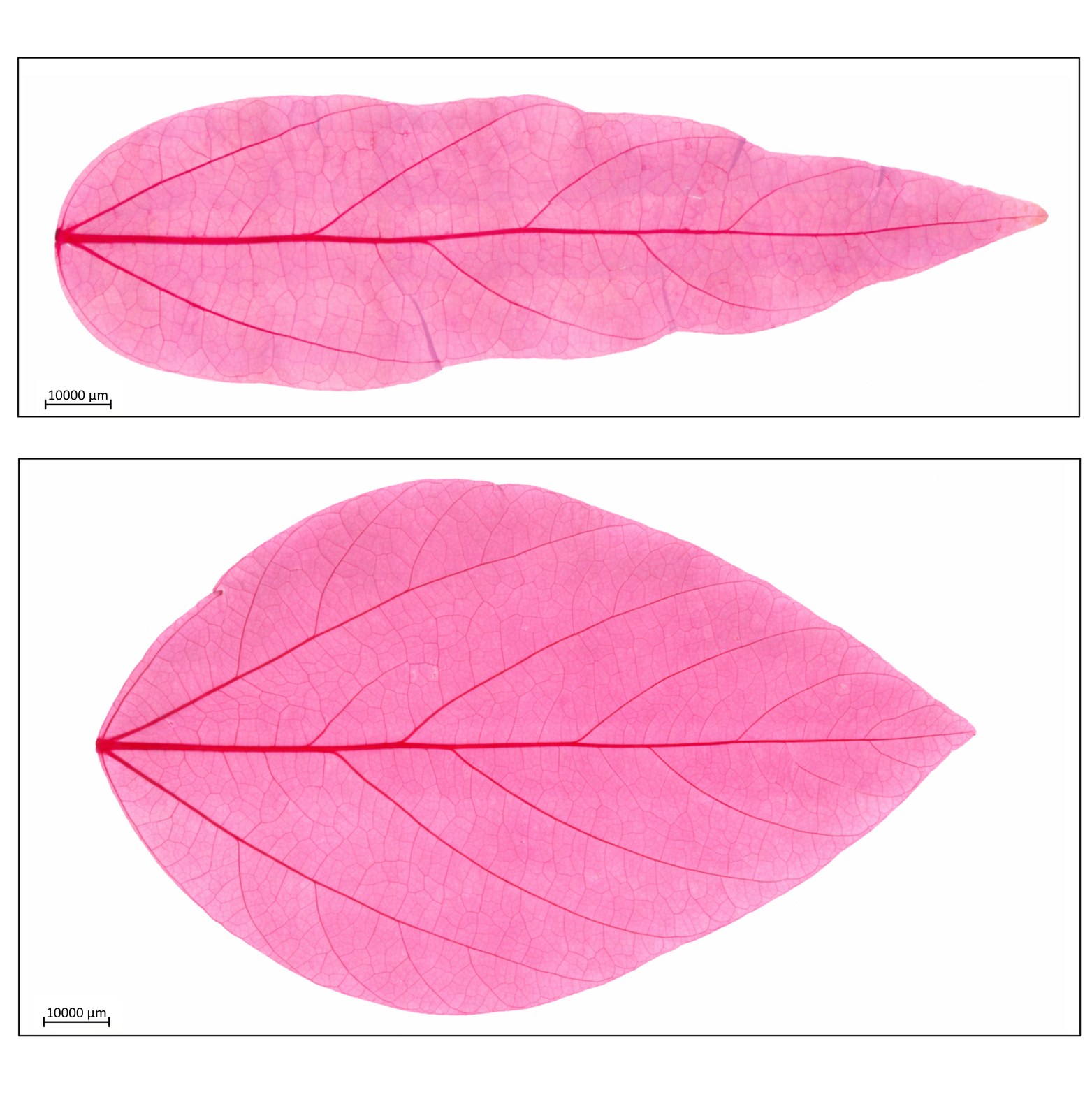








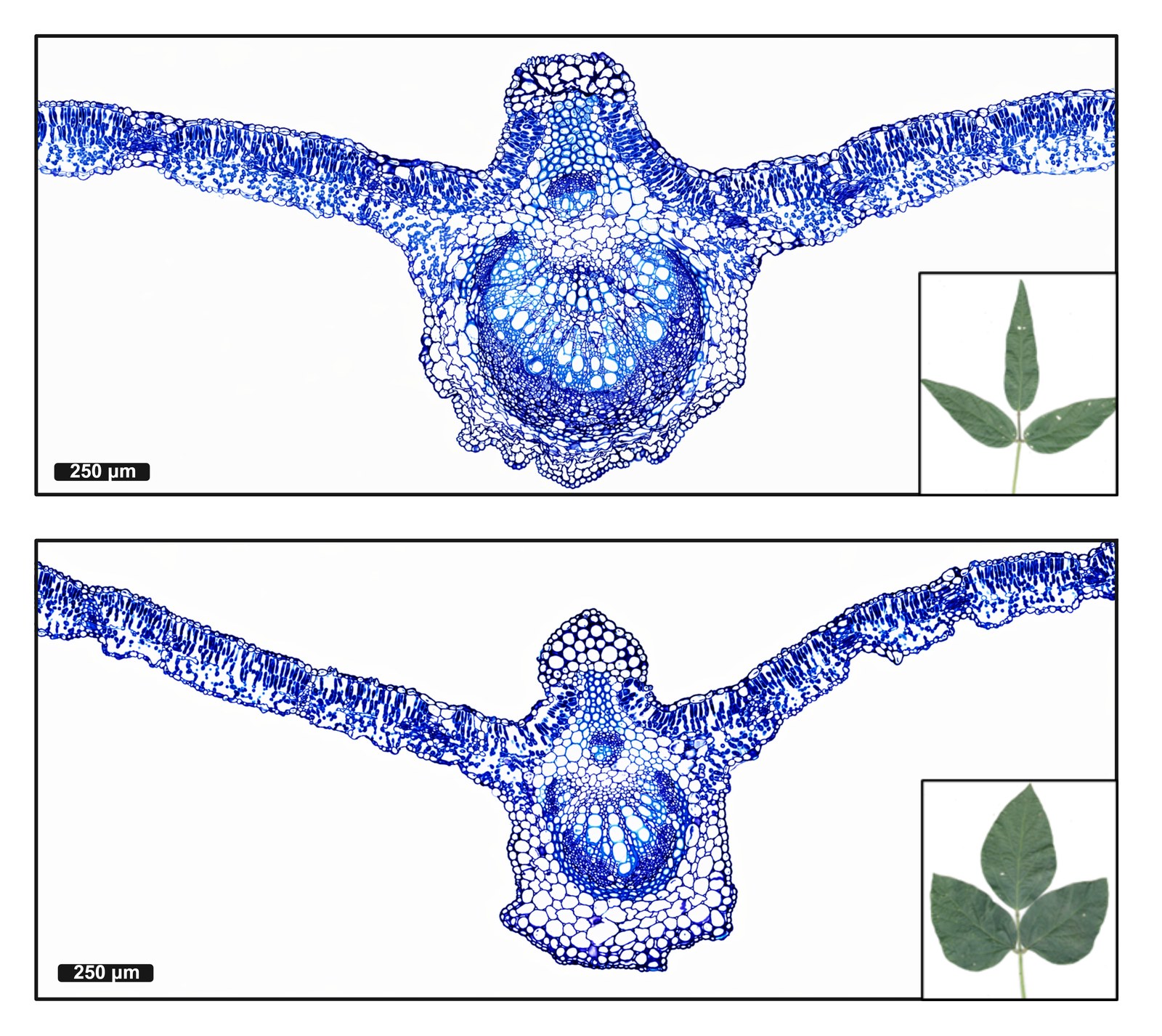










Academic Background
Education
Agronomy and Crop Sciences
Virginia Tech
Plant Breeding and Genetics
Tribhuvan University
Plant Breeding and Genetics
Tribhuvan University
Research Positions
Research Scientist
University of Illinois
Postdoctoral Fellow
University of Minnesota
Graduate Research Assistant
Virginia Tech
Selected Awards
RIPE Project Ambassador
ASPB Plantae's Network Champion
Charles I. Rich Fellowship
Virginia Tech
Phyllis G. & Reginald H. Nelson IV
Tuition Scholarship
Contact
Office
Institute for Genomic Biology
University of Illinois
Urbana-Champaign, IL 61801